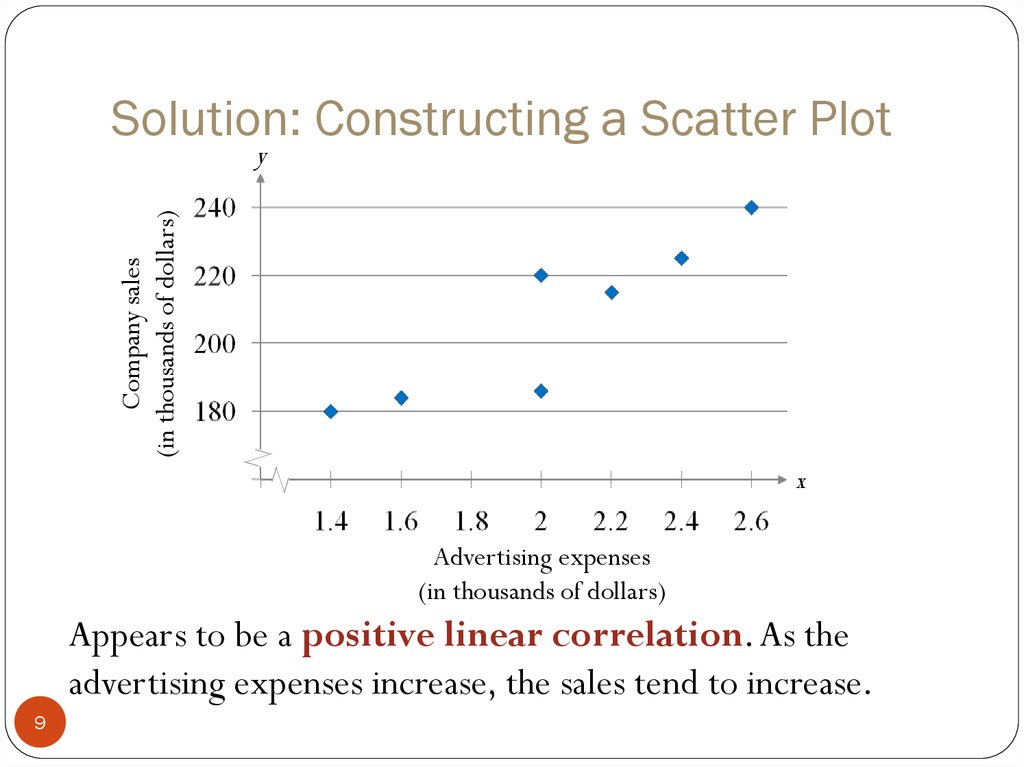
Other tools to characterize cell-secreted products lack the quantitative resolution, throughput, and multiplexing of flow cytometry and do not directly link secretions to transcriptomic information. 
However, transcript levels do not account for various downstream processes after transcription, including RNA splicing, translation, post-translational modifications, enzymatic cleavage, or even storage of secreted proteins in secretory vesicles prior to secretion in response to a stimulus. Similarly, transcripts associated with secreted proteins may sometimes correlate to secretion 1, 2, 3. Furthermore, the presence of secreted proteins on or within the cell does not necessarily indicate that these proteins would have been secreted. Downsides of this permeabilization and fixation process include the loss of cell viability and loss of other intracellular molecules, such as mRNA, which limits downstream transcriptomic analysis. Forming pores in the cell is necessary for fluorescently-labeled antibodies to penetrate the intracellular space and bind to chemically fixed proteins. 
Intracellular proteins, including secreted proteins, can be analyzed, e.g., using intracellular cytokine staining, but through a destructive process that involves permeabilization and fixation of the cell. One possible method is by combining flow cytometry with intracellular staining to quantify intracellular production of secreted proteins in the context of other markers. Standard single-cell analysis tools cannot simultaneously assess external cellular information (e.g., secreted proteins) with cell surface and/or intracellular information. Linking these functional changes in the capacity for secretion of immunoglobulins to genetic/phenotypic profiles at the single-cell level can uncover the population heterogeneity and potential new cell states. In this process, B cells differentiate, undergoing significant phenotypic, morphological, and genetic changes. For example, one of the main roles of B cells is to respond to antigens with the production and secretion of large quantities of immunoglobulins targeting antigen epitopes. Organisms critically depend on the proteins or other factors which cells secrete into their environment that can act locally in a paracrine manner or systemically. Altogether, this method links quantity of secretion with single-cell sequencing (SEC-seq) and enables researchers to fully explore the links between genome and function, laying the foundation for discoveries in immunology, stem cell biology, and beyond. By using oligonucleotide-labeled antibodies we find that upregulation of pathways for protein localization to the endoplasmic reticulum and mitochondrial oxidative phosphorylation are most associated with high IgG secretion, and uncover surrogate plasma cell surface markers (e.g., CD59) defined by the ability to secrete IgG. Measurements using flow cytometry and imaging flow cytometry corroborate the association between IgG secretion and CD38/CD138. By accumulating secretions close to secreting cells held within cavity-containing hydrogel nanovials, we demonstrate workflows to analyze the amount of IgG secreted from single human B cells and link this information to surface markers and transcriptomes from the same cells. The argument method allows you to select between "circle" (default), "square", "ellipse", "number", "shade", "pie", and "color".The secreted products of cells drive many functions in vivo however, methods to link this functional information to surface markers and transcriptomes have been lacking. You can use the colorRampPalette function to generate color spectra. Stars = TRUE, # If TRUE, adds significance level with starsĬi = TRUE) # If TRUE, adds confidence intervals Jiggle = FALSE, # If TRUE, data points are jittered Lm = FALSE, # If TRUE, plots linear fit rather than the LOESS (smoothed) fitĬor = TRUE, # If TRUE, reports correlations Method = "pearson", # Correlation method (also "spearman" or "kendall") Scale = FALSE, # If TRUE, scales the correlation text fontĭensity = TRUE, # If TRUE, adds density plots and histogramsĮllipses = TRUE, # If TRUE, draws ellipses Smooth = TRUE, # If TRUE, draws loess smooths The pairs.panel function is an extension of the pairs function that allows you to easily add regression lines, histograms, confidence intervals, … and customize several additional arguments. The package pysch provides two interesting functions to create correlation plots in R.
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
